General Practitioners in parisian departements¶
[1]:
from pynsee.sirene import get_dimension_list, search_sirene
from pynsee.metadata import get_activity_list
from pynsee.geodata import get_geodata_list, get_geodata
from pynsee.localdata import get_geo_list
import difflib
import geopandas as gpd
import numpy
import pandas as pd
from pandas.api.types import CategoricalDtype
import matplotlib.cm as cm
import matplotlib.pyplot as plt
import descartes
import re
from shapely.geometry import Polygon, MultiPolygon
from matplotlib.offsetbox import AnchoredText
[2]:
import logging
import sys
logging.basicConfig(stream=sys.stdout,
level=logging.INFO,
format="%(message)s")
[4]:
get_dimension_list().to_csv("sirene_dimensions.csv")
Existing environment variables used, instead of locally saved credentials
1 - Getting data: 100%|██████████| 1/1 [00:00<00:00, 102.94it/s]
Data saved: /home/onyxia/.cache/pynsee/pynsee/76f10aaaadf5a625061805b71f0bf0db
This function renders package's internal data
This function may return personal data, please check and comply with the legal framework relating to personal data protection !
[5]:
# get activity list
naf5 = get_activity_list('NAF5')
naf5.to_csv("naf.csv")
naf5[naf5["NAF5"].str.contains("^86.2")]
[5]:
| A10 | A129 | A17 | A21 | A38 | A5 | A64 | A88 | NAF1 | NAF2 | NAF3 | NAF4 | NAF5 | TITLE_NAF5_40CH_FR | TITLE_NAF5_65CH_FR | TITLE_NAF5_FR | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 664 | OQ | Q86Z | OQ | Q | QA | OQ | QA0 | 86 | Q | 86 | 86.2 | 86.21 | 86.21Z | Activité des médecins généralistes | Activité des médecins généralistes | Activité des médecins généralistes |
| 665 | OQ | Q86Z | OQ | Q | QA | OQ | QA0 | 86 | Q | 86 | 86.2 | 86.22 | 86.22A | Act. radiodiagnostic et de radiothérapie | Activités de radiodiagnostic et de radiothérapie | Activités de radiodiagnostic et de radiothérapie |
| 666 | OQ | Q86Z | OQ | Q | QA | OQ | QA0 | 86 | Q | 86 | 86.2 | 86.22 | 86.22B | Activités chirurgicales | Activités chirurgicales | Activités chirurgicales |
| 667 | OQ | Q86Z | OQ | Q | QA | OQ | QA0 | 86 | Q | 86 | 86.2 | 86.22 | 86.22C | Autre activité des médecins spécialistes | Autres activités des médecins spécialistes | Autres activités des médecins spécialistes |
| 668 | OQ | Q86Z | OQ | Q | QA | OQ | QA0 | 86 | Q | 86 | 86.2 | 86.23 | 86.23Z | Pratique dentaire | Pratique dentaire | Pratique dentaire |
[6]:
# departements list of ile de france
list_dep = ["92", "93", "75", "94"] # + ["77", "78", "95", "91"]
string_dep = "|".join([dep + "*" for dep in list_dep])
string_dep
[6]:
'92*|93*|75*|94*'
[7]:
# search data on doctors
doctors = search_sirene(variable = ["activitePrincipaleEtablissement", "codePostalEtablissement"],
pattern = ['86.21Z*', string_dep],
number = 100000)
# '86.21Z' médecin généraliste
doctors = doctors[doctors["activitePrincipaleEtablissement"].str.contains("^86.21Z")].reset_index(drop=True)
1 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 212.20it/s]
2/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 224.17it/s]
3/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 207.62it/s]
4/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 217.40it/s]
5/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 225.08it/s]
6/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 222.84it/s]
7/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 234.79it/s]
8/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 204.70it/s]
9/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 216.98it/s]
10/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 220.54it/s]
11/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 220.24it/s]
12/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 232.69it/s]
13/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 236.86it/s]
14/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 230.69it/s]
15/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 234.82it/s]
16/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 240.52it/s]
17/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 238.77it/s]
18/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 241.84it/s]
19/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 226.83it/s]
20/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 222.08it/s]
21/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 238.45it/s]
22/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 241.57it/s]
23/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 245.45it/s]
24/100 - Getting data: 100%|██████████| 1000/1000 [00:03<00:00, 252.45it/s]
25/100 - Getting data: 100%|██████████| 1000/1000 [00:04<00:00, 237.15it/s]
26/100 - Getting data: 100%|██████████| 1000/1000 [00:03<00:00, 261.68it/s]
27/100 - Getting data: 100%|██████████| 1000/1000 [00:03<00:00, 258.79it/s]
28/100 - Getting data: 100%|██████████| 1000/1000 [00:03<00:00, 257.36it/s]
29/100 - Getting data: 100%|██████████| 175/175 [00:00<00:00, 255.57it/s]
Data saved: /home/onyxia/.cache/pynsee/pynsee/6c0f248de2346ddc6cb0d424b41d8f40
[8]:
# filter
list_pattern = ["RADIOLOGIE", "IMAGERIE", "GYNECO", "DENTAIRE", "ANESTHES", 'P.M.I', 'INFANTILE', 'CARDIOLOG',
"MAXILLO", "HOSPITALIER", "OPHTALMO", "DERMATOLOG", 'PLANIFICATION', 'MAIRIE', 'THORACIQUE',
"TOMODENSITOM", "URGENCE", "NEPHROLOG", "PRAXEA", 'VACCINATION', 'SCOLAIRE', 'CANCER', 'STOMATOLO',
"URODYNAMIQ", "CARDIORIS", "CHIRPLASTIQUE", "NINAMAX", "CICOF", 'MATERNELLE', 'DIALYSE',
'RADIOTHERAPIE', 'RHUMATOLOG', 'DENTISTE', 'ECHOGRAPH', 'PATHOLOG', 'CHIRURGIE', 'ESTHETIQ',
'CYTOLOG', 'ORTHOPED']
pattern = r"|".join(list_pattern)
[9]:
# clean up the data not related to general practitioners but to other health activities
doctors = doctors[~doctors["denominationUniteLegale"].str.contains(pattern, regex=True, na=False)].reset_index(drop=True)
doctors = doctors[~doctors["enseigne1Etablissement"].str.contains(pattern, regex=True, na=False)].reset_index(drop=True)
# exclude association and administration
doctors = doctors[~doctors["categorieJuridiqueUniteLegale"].isin(["9220", "7220"])].reset_index(drop=True)
doctors = doctors.sort_values(["categorieJuridiqueUniteLegale"])
doctors.to_csv("doctors.csv")
list_denomination_uniteLegale = doctors.denominationUniteLegale.unique()
[10]:
# get geographical data list
geodata_list = get_geodata_list()
# get communes geographical limits
com = get_geodata('ADMINEXPRESS-COG-CARTO.LATEST:commune')
# get arrondissement (subdivision of departement) geographical limits
arr = get_geodata('ADMINEXPRESS-COG-CARTO.LATEST:arrondissement')
Previously saved data has been used:
/home/onyxia/.cache/pynsee/pynsee/2ac583be54e866b2b1b30fb0113c1fd0
Set update=True to get the most up-to-date data
100%|██████████| 99/99 [00:33<00:00, 2.97it/s]
Data saved: /home/onyxia/.cache/pynsee/pynsee/a4f589755961ab3aa6bdff99605334d1
Data saved: /home/onyxia/.cache/pynsee/pynsee/1b27559c061e4bdaa0bd2966f4605287
[11]:
# get arrondissement municipal geographical limits
arrmun = get_geodata('ADMINEXPRESS-COG-CARTO.LATEST:arrondissement_municipal')
arrmun = arrmun[arrmun.insee_com.str.contains('^75')].reset_index(drop=True)
arrmun["nom_m"] = arrmun["nom_m"].apply(lambda x: re.sub(r"ER|E", "", re.sub(" ARRONDISSEMENT", "", x)))
arrmun = arrmun.rename(columns={"geometry" : "geometry2",
"population" : "population2",
"nom_m" : "nom_m2"})
arrmun = arrmun[["nom_m2", "geometry2", "population2"]]
Data saved: /home/onyxia/.cache/pynsee/pynsee/94d0586636b45c9d8cdd466da4a3f3b8
[12]:
def _replace_nan(x, y):
if (x is None) or (numpy.isnan(x)):
return y
else:
return x
[13]:
doctors
[13]:
| siren | nic | siret | dateDebut | dateCreationEtablissement | dateCreationUniteLegale | dateFin | denominationUniteLegale | nomUniteLegale | prenomUsuelUniteLegale | ... | libellePaysEtrangerEtablissement | complementAdresse2Etablissement | indiceRepetition2Etablissement | libelleCommuneEtranger2Etablissement | distributionSpeciale2Etablissement | codeCedex2Etablissement | libelleCedex2Etablissement | codePaysEtranger2Etablissement | libellePaysEtranger2Etablissement | enseigne3Etablissement | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 5267 | 445161631 | 00038 | 44516163100038 | 2014-11-01 | 2014-11-01 | 2002-12-31 | None | None | BACH | DANIEL | ... | None | None | None | None | None | None | None | None | None | None |
| 6477 | 518143185 | 00016 | 51814318500016 | 2009-11-02 | 2009-11-02 | 2009-11-02 | None | None | RICCI | AXEL | ... | None | None | None | None | None | None | None | None | None | None |
| 6475 | 518032172 | 00026 | 51803217200026 | 2013-06-01 | 2013-06-01 | 2009-05-02 | None | None | CHENIOUR | SOUMAYA | ... | None | None | None | None | None | None | None | None | None | None |
| 6474 | 518023916 | 00027 | 51802391600027 | 2012-01-02 | 2012-01-02 | 2009-10-31 | None | None | BIBAS | AURORE | ... | None | None | None | None | None | None | None | None | None | None |
| 6473 | 518001169 | 00029 | 51800116900029 | 2015-12-02 | 2015-12-02 | 2009-06-05 | None | None | SIDHOUM ANNAB | SAMIRA | ... | None | None | None | None | None | None | None | None | None | None |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 6916 | 775665805 | 00468 | 77566580500468 | 2008-01-01 | 1996-12-25 | 1900-01-01 | None | COMPAGNIE FILLES CHARITE ST VINCENT PAUL | None | None | ... | None | None | None | None | None | None | None | None | None | None |
| 6928 | 775683006 | 00362 | 77568300600362 | 2017-06-21 | 2017-06-21 | 1900-01-01 | None | FONDATION SANTE DES ETUDIANTS DE FRANCE | None | None | ... | None | None | None | None | None | None | None | None | None | None |
| 6915 | 775663883 | 00020 | 77566388300020 | 2008-01-01 | 1900-01-01 | 1900-01-01 | None | INSTITUT ARTHUR VERNES | None | None | ... | None | None | None | None | None | None | None | None | None | None |
| 292 | 313711038 | 00015 | 31371103800015 | 2008-01-01 | 1978-01-01 | 1978-01-01 | None | FONDATION HAHNEMANN | None | None | ... | None | None | None | None | None | None | None | None | None | None |
| 221 | 310227749 | 00010 | 31022774900010 | 2008-01-01 | 1900-01-01 | 1900-01-01 | None | CENTRE DE SOINS | None | None | ... | None | None | None | None | None | None | None | None | None | None |
10536 rows × 91 columns
[14]:
# count general practioners by commun
hdLegalStatus = doctors.groupby(['categorieJuridiqueUniteLegale'], dropna=True)['effectifsMinEtablissement'].agg(['mean', "count"])
hdLegalStatus["meanHeadcountLegalStatus"] = hdLegalStatus["mean"].apply(lambda x: _replace_nan(x, 1))
hdLegalStatus["categorieJuridiqueUniteLegale"] = hdLegalStatus.index
hdLegalStatus.loc[hdLegalStatus["categorieJuridiqueUniteLegale"] == "1000", "meanHeadcountLegalStatus"] = 1
hdLegalStatus = hdLegalStatus.reset_index(drop=True)
doctors = doctors.merge(hdLegalStatus, how = "left", on = "categorieJuridiqueUniteLegale").reset_index(drop=True)
for r in doctors.index:
hdMin = doctors.loc[r, "effectifsMinEtablissement"]
if (hdMin is None) or (numpy.isnan(hdMin)):
doctors.loc[r, "headcount"] = doctors.loc[r, "meanHeadcountLegalStatus"]
else:
doctors.loc[r, "headcount"] = hdMin
[15]:
# duplicates cleaning
# normalization to avoid double counting
doctors["headcountFinal"] = doctors["headcount"] / doctors.groupby("siren", dropna=True)["headcount"].transform("sum")
#visual check duplicates
dupDoctors = doctors[doctors["siren"].duplicated(keep=False)].reset_index(drop=True)
dupDoctors.to_csv("dupDoctors.csv")
list_adress_var = ["numeroVoieEtablissement", "typeVoieEtablissementLibelle",
"libelleVoieEtablissement", "codePostalEtablissement", "libelleCommuneEtablissement"]
addressDupDoctors = doctors[doctors[list_adress_var].duplicated(keep=False)].reset_index(drop=True)
addressDupDoctors = addressDupDoctors.sort_values(list_adress_var)
addressDupDoctors.to_csv("addressDupDoctors.csv")
[16]:
doctors.sort_values(["libelleCommuneEtablissement"]).to_csv("doctorSortedCom.csv")
[17]:
# count general practioners by commune
doctorsCom = doctors.groupby(['libelleCommuneEtablissement', 'codePostalEtablissement'])['headcountFinal'].agg(['sum'])
doctorsCom = doctorsCom.rename(columns = {"sum" : "headcount"})
doctorsCom
[17]:
| headcount | ||
|---|---|---|
| libelleCommuneEtablissement | codePostalEtablissement | |
| ABLON-SUR-SEINE | 94480 | 2.000000 |
| ALFORTVILLE | 94140 | 31.250000 |
| ANTONY | 92160 | 144.601926 |
| ARCUEIL | 94110 | 15.500000 |
| ASNIERES-SUR-SEINE | 92600 | 63.750000 |
| ... | ... | ... |
| VILLEPINTE | 93420 | 24.000000 |
| VILLETANEUSE | 93430 | 4.000000 |
| VILLIERS-SUR-MARNE | 94350 | 15.000000 |
| VINCENNES | 94300 | 76.500000 |
| VITRY-SUR-SEINE | 94400 | 86.250000 |
147 rows × 1 columns
[18]:
doctorsCom["libelleCommuneEtablissement"] = doctorsCom.index.get_level_values("libelleCommuneEtablissement")
doctorsCom["codePostalEtablissement"] = doctorsCom.index.get_level_values("codePostalEtablissement")
doctorsCom = doctorsCom.reset_index(drop=True)
[19]:
import os, sys
communes = get_geo_list("communes")
def _find_commune_identifier(com, dep, communes=communes):
try:
list_com = [c.upper() for c in communes.TITLE.to_list()]
communes["TITLE"] = list_com
communes = communes[communes["CODE_DEP"] == dep].reset_index(drop=True)
match = difflib.get_close_matches(com.upper(), list_com)[0]
cog = communes[communes["TITLE"] == match].reset_index(drop=True)
cog = cog.loc[0, "CODE"]
except:
cog = None
return cog
for r in doctorsCom.index:
dep = doctorsCom.loc[r, "codePostalEtablissement"][:2]
comName = doctorsCom.loc[r, "libelleCommuneEtablissement"]
doctorsCom.loc[r, "insee_com"] = _find_commune_identifier(comName, dep)
[20]:
comDep = com[com["insee_dep" ].isin(list_dep)].reset_index(drop=True)
doctorsCom2 = comDep.merge(doctorsCom, how = "left", on = "insee_com")
[21]:
doctorsCom2["headcount"] = doctorsCom2["headcount"].apply(lambda x: _replace_nan(x, 0))
[22]:
doctorsCom2["headcount"] = doctorsCom2["headcount"].apply(lambda x: _replace_nan(x, 0))
doctorsCom2["ratio"] = 1000 * doctorsCom2["headcount"] / doctorsCom2["population"]
[23]:
doctorsCom2 = doctorsCom2.reset_index(drop=True)
doctorsCom4 = doctorsCom2[doctorsCom2.columns].merge(arrmun, left_on ="libelleCommuneEtablissement", right_on="nom_m2", how="left")
for i in doctorsCom4.index:
if type(doctorsCom4.loc[i, 'geometry2']) in [MultiPolygon, Polygon]:
doctorsCom4.loc[i, 'geometry'] = doctorsCom4.loc[i, 'geometry2']
doctorsCom4.loc[i, "ratio"] = 1000 * doctorsCom4.loc[i,"headcount"] / doctorsCom4.loc[i,"population2"]
del doctorsCom4["geometry2"]
doctorsCom2Map = gpd.GeoDataFrame(doctorsCom4)
[24]:
doctorsCom2Map = gpd.GeoDataFrame(doctorsCom4)
[25]:
# compute density ranges of GP
doctorsCom2Map.loc[doctorsCom2Map.ratio < 1, 'range'] = "< 0.2"
doctorsCom2Map.loc[doctorsCom2Map.ratio >= 5, 'range'] = "> 5"
density_ranges = [0.2, 0.6, 1, 1.5, 2, 2.5, 3, 3.5, 5]
list_ranges = []
list_ranges.append( "< 0.2")
for i in range(len(density_ranges)-1):
min_range = density_ranges[i]
max_range = density_ranges[i+1]
range_string = "[{}, {}[".format(min_range, max_range)
rows = (doctorsCom2Map.ratio >= min_range) & (doctorsCom2Map.ratio < max_range)
doctorsCom2Map.loc[rows, 'range'] = range_string
list_ranges.append(range_string)
list_ranges.append("> 5")
doctorsCom2Map['range2'] = doctorsCom2Map['range'].astype(CategoricalDtype(categories=list_ranges, ordered=True))
[26]:
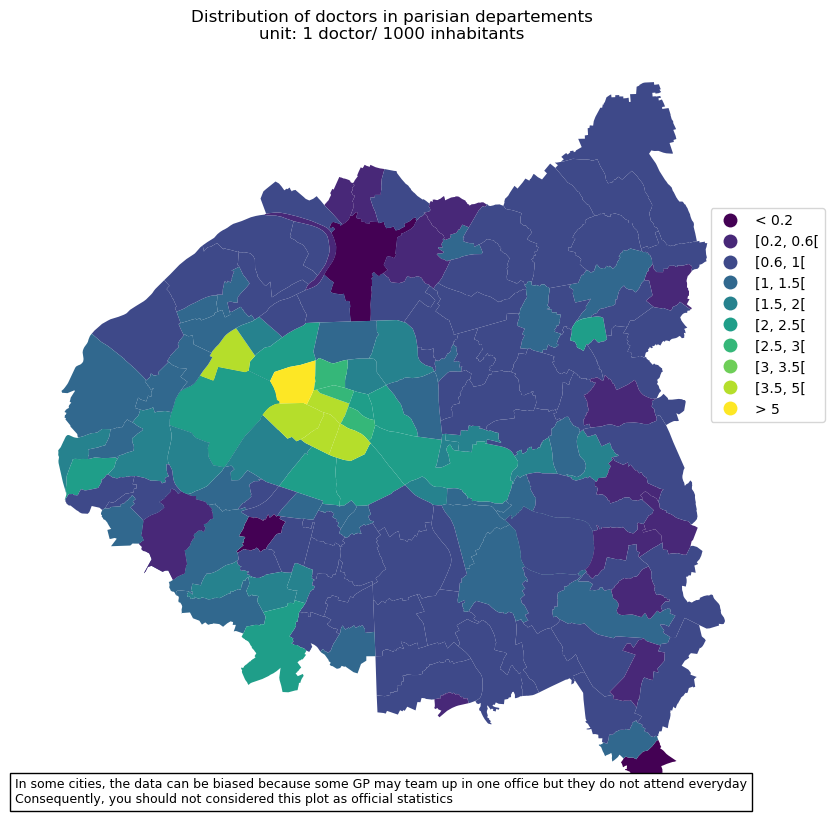
# make plot
txt = 'In some cities, the data can be biased because some GP may team up in one office but they do not attend everyday'
txt += '\nConsequently, you should not considered this plot as official statistics'
fig, ax = plt.subplots(1,1,figsize=[10,10])
doctorsCom2Map.plot(column='range2', cmap=cm.viridis,
legend=True, ax=ax,
legend_kwds={'bbox_to_anchor': (1.1, 0.8)})
ax.set_axis_off()
ax.set(title='Distribution of doctors in parisian departements\nunit: 1 doctor/ 1000 inhabitants')
at = AnchoredText(txt, prop=dict(size=9), frameon=True, loc=4)
ax.add_artist(at)
plt.show()
[ ]: